Capture-Seq makes studying evolutionary relationships across broad taxonomic groups easier, faster, and more cost-effective than ever. Use Capture-Seq data to answer research questions for:
We deliver custom solutions for any species or taxonomic group, using public resources or even generating new ones. Ask about our strategies for never-been-done questions.
Our full-service Capture-Seq pipeline is optimized for challenging sample types like degraded, low-input museum and herbarium specimens. Our industry-leading automation ensures robust, reliable results whether you have dozens of samples at once or tens of thousands of samples across several years.
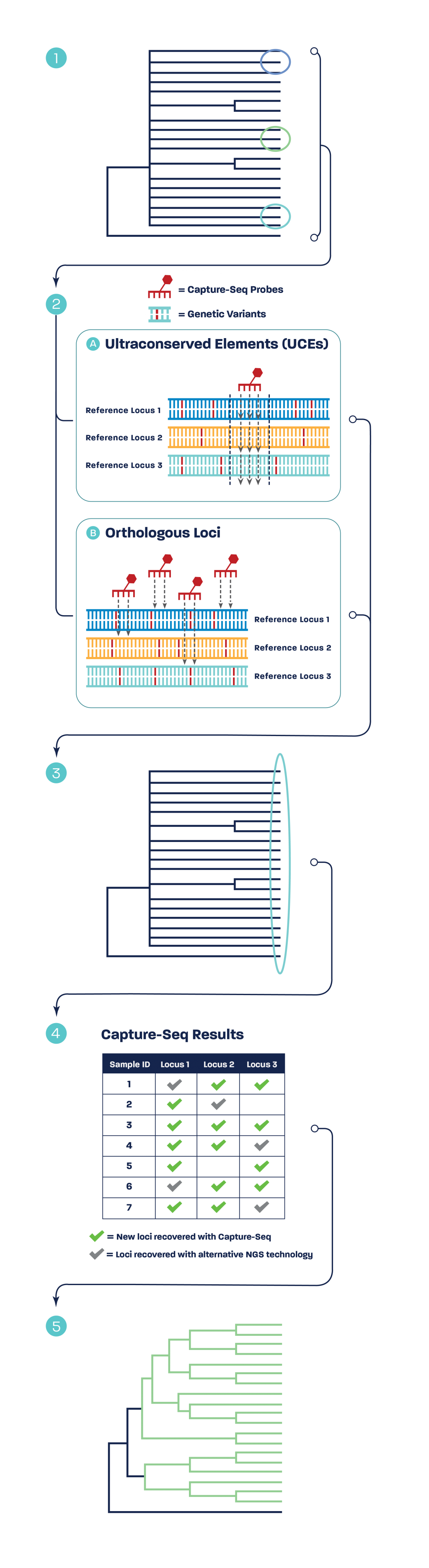
Capture-Seq’s technology uses targeted probe hybridization enrichment to selectively sequence and assemble loci shared between as few as one species to an entire taxonomic clade. By focusing only on informative regions of the genome, Capture-Seq is more cost-effective than whole-genome sequencing.
Capture-Seq datasets are more complete and more tolerant of degraded samples than restriction enzyme genotyping-by-sequencing (GBS), yielding consistently superior results and easier analyses. Mitochondrial and chloroplast data can be obtained as a by-product of nuclear Capture-Seq data, or by targeting either genome directly.
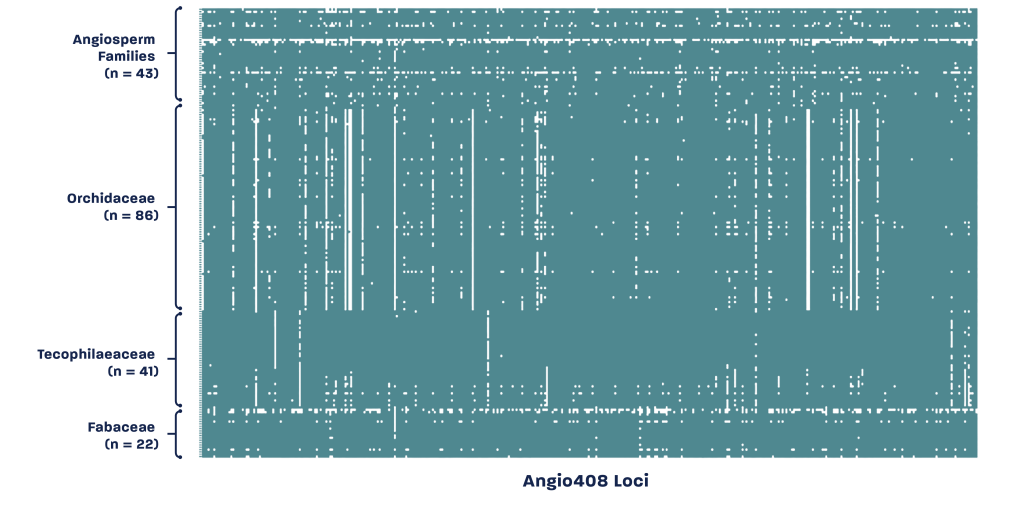
The Capture-Seq Angiosperms-408 (Angio408) probe set captures single-copy nuclear loci for phylogenetic studies across the angiosperms. With an emphasis on recovering full-length sequencing data from 408 expertly curated genomic regions, the Angio408 probe set delivers a one-size-fits-all phylogenetic tool designed to effectively resolve relationships across all flowering plant clades.
The Angio408 probe set was developed at Rapid Genomics in collaboration with Gordon Burleigh (University of Florida).
UCE sequencing is a specialized application of the Capture-Seq platform, where probes target highly conserved regions in genomes across distant evolutionary taxa for sequencing and comparative analysis.
The selective sequencing of UCE’s and their flanking regions have powerful applications in evolutionary and population genetics studies.
In addition to full-service UCE solutions from sample processing to sequencing results, Rapid Genomics offers custom bioinformatics packages using the Phyluce pipeline and other analytical pipelines.
LEARN MORE ABOUT BIOINFORMATICS
Designed from 2,628 baits targeting 1,314 conserved loci identified among Acanthomorphs (fishes).
Described as part of Alfaro et al. 2013.
Designed from 31,829 baits targeting 2,590 conserved loci and exons identified among Anthozoans (sea anemones and corals).
Described as part of Quattrini et al. 2017.
Designed from 13,674 baits targeting 1,172 conserved loci identified among Coleoptera (beetles).
Described as part of Faircloth 2017.
Designed from 40,207 baits targeting 2,731 conserved loci identified among Hemiptera (true bugs).
Described as part of Faircloth 2017.
An expanded version of the Hymenoptera 1.5Kv1 probe set. Designed from 31,829 baits targeting 2,590 conserved loci identified among Hymenoptera (wasps, bees, ants, sawflies).
Described as part of Branstetter et al. 2017.
Designed from 6,737 baits targeting 2,708 conserved loci identified among Ostariophysans (fishes).
Described as part of Faircloth et al. 2020.
Designed from 5,472 baits targeting 5,060 conserved loci among Tetrapods (birds, mammals, reptiles, amphibians).
Described as part of Faircloth et al. 2012.
Designed from 2,001 baits targeting 500 conserved loci identified among Actinopterygians (ray-finned fishes).
Described as part of Faircloth et al. 2013.
Designed from 14,799 baits targeting 1,120 conserved loci identified among arachnids (scorpions, ticks, mites, harvestmen, and solifuges)
Described as part of Faircloth 2017.
Designed from 31,328 baits targeting 2,711 conserved loci identified among Diptera (true flies).
Described as part of Faircloth 2017.
Designed from 2,749 baits targeting 1,510 conserved loci identified among Hymenoptera (wasps, bees, ants, sawflies).
Described as part of Faircloth et al. 2015.
A subset of the Hymenoptera 2.5Kv2 probe set. Designed from 9,898 baits targeting 2,524 conserved loci and 16 exons specific to ants.
Described as part of Branstetter et al. 2017.
Designed from 2,560 baits targeting 2,386 conserved loci identified among Tetrapods (birds, mammals, reptiles, amphibians).
Described as part of Faircloth et al. 2012.