Stop wasting, start saving
Every genome contains repetitive elements that are not suitable for WGS analysis, translating to wasted resources on sequencing. This is particularly true in low-pass genome skimming and imputation, where efficiently generating <1x sequencing depth is essential to cost-effective use in commercial agriculture
GD-Seq (repetitive element genome depletion sequencing) maximizes your efficiency sequencing resources, improving data in low-copy, non-repetitive genomic regions for easier imputation and breeding selections.
GD-Seq minimizes waste by removing repetitive DNA elements and boosting coverage of the most informative, low copy genomic regions.
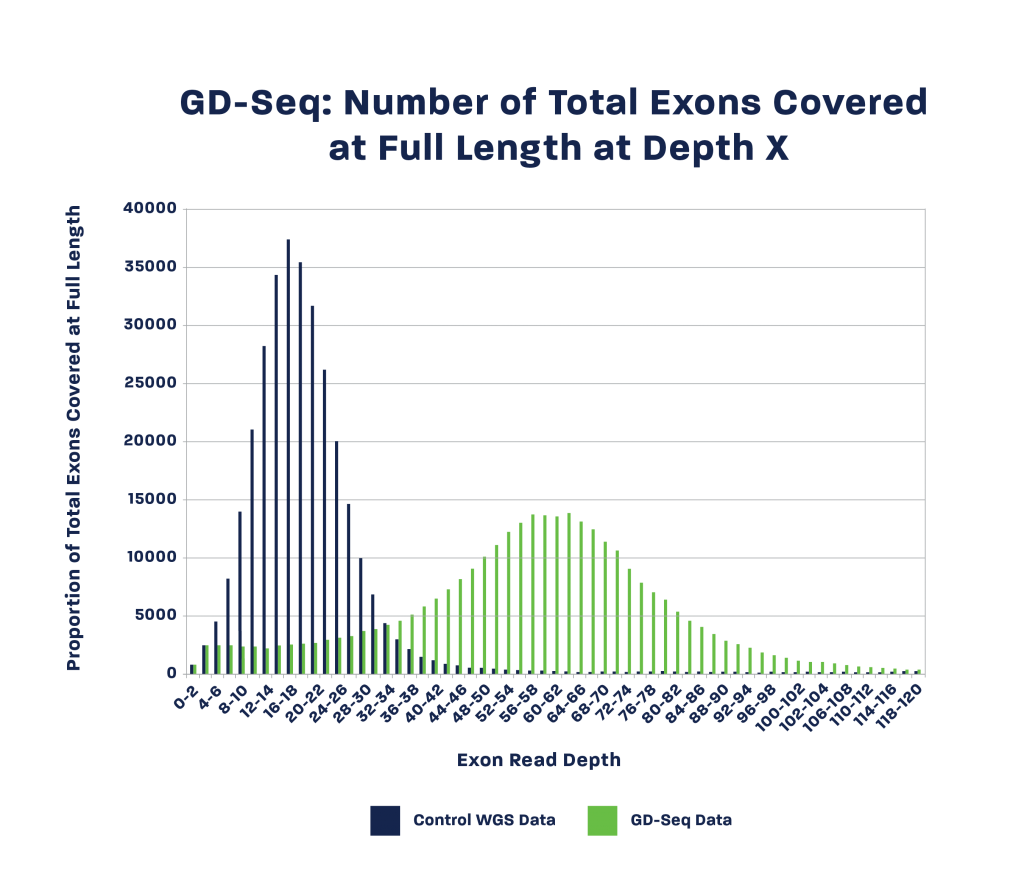
These advances over WGS technology improve exon coverage by up to 500% at equivalent sequencing resources, delivering a new level of cost-efficiency at any scale.
Maize samples were processed using GD-Seq and traditional whole-genome sequencing protocols. Both datasets were generated using equal sequencing efforts and analyzed for read depth in all exons of the B73 reference genome. GD-Seq results recovered 90% of all exons and sequencing depth improved by 300% – 500%, suitable for discovery and genome skimming applications.
In addition to end-to-end GD-Seq solutions, we provide specialized data analysis and bioinformatics support, including but not limited to: